Hierarchically-Organized Latent Modules for
Exploratory Search in Morphogenetic Systems
Mayalen Etcheverry, Clément Moulin-Frier, Pierre-Yves Oudeyer
June 2020
A Morphogenetic System: Lenia
We use Lenia cellular automaton, a continuous generalization to Conway’s Game of Life, as environment of study.
Lenia can generate a wide range of complex behaviors and life-like structures, as testifies the collection of “species” that have been manually identified and classified by Lenia’s crator Bert Chan.
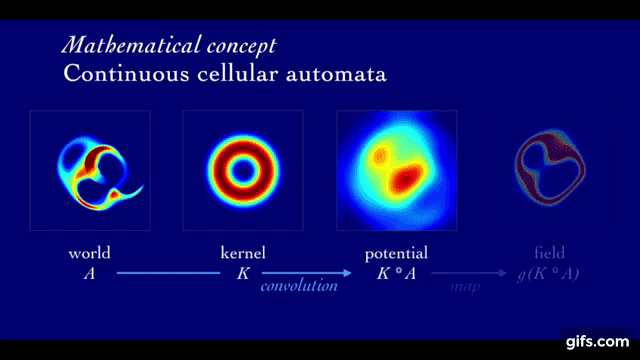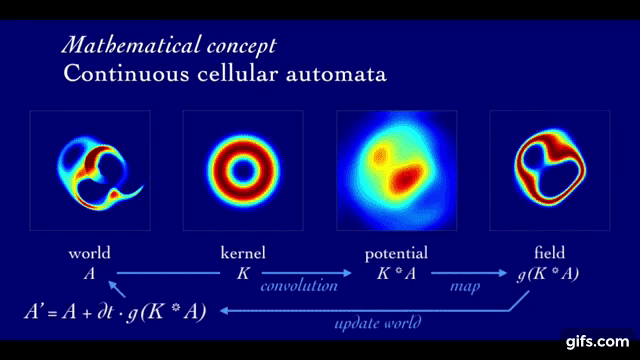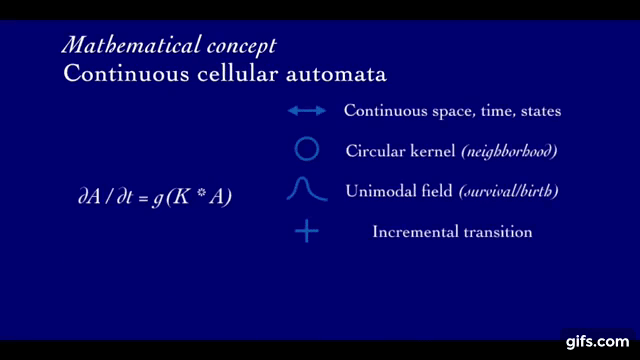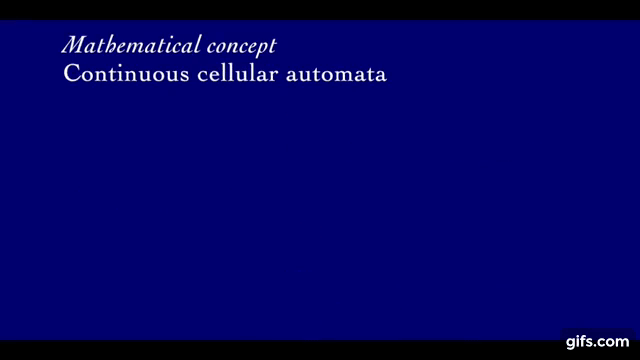
A rollout in Lenia is defined by the initial state (At=1, seed of the system) as well as its local update rule that is iteratively applied to evolve the state of the system through time (At=1 → … → At=T).
In this paper, an artificial agent is given the task to explore Lenia with a budget of N=5000 runs, where, for each run the agent must select the set of initial parameters θ ∈ Θ that will determine the system rollout. We aim to build an artificial “discovery assistant” that would automatically assist a human end-user in its exploration of a complex dynamical system like Lenia.
Proposed Approach: IMGEP-HOLMES
In IMGEP-HOLMES, a hierarchy of embedding networks is actively constructed by the exploring agent to represent the different niches of patterns discovered during the exploration loop. Below, we provide a visualisation tool to browse through the discovered niches of patterns within the tree-structured representation. We can observe that IMGEP-HOLMES unsupervisedly learns to separate visually distinct categories of patterns into the different behavioral characterization spaces, allowing to discover diverse patterns within each of them.
BC 0:
Patterns are visualized with 3D PCA reduction of the original 16D BC space. You can rotate (left click), pan (right click) and scroll (mouse wheel) through the patterns.
This is an example of visualisation tool that could be used for integrating a human evaluator in the loop.
With only few “clicks”, the human can browse through the different modules and assign preferences scores in order to guide the exploration process.
Example of Interesting Niche of Patterns Autonomously Discovered by IMGEP-HOLMES
Continuous cellular automata are of particular interest to better understand the chemical origin of Life. Indeed, a major mystery in understanding life’s beginning is the possibility that complex ordered patterns (closed systems) could poped to existence out of an initially disordered state, referred to as primmordial soup. In the extended Lenia paper, many parallels are done with the artificial phenomenon observed in Lenia:
The gradual emergence of several important phenomena in Lenia is reminiscent of the origin of life. Cell individuality and self-replication are among the hallmarks of life on Earth, each has abiotic origins. Individuality originated from lipid membranes that were formed spontaneously by hydrophobic molecules in the primordial soup, separate the outside world from an area where specific chemical reactions can occur, and protect such an area from physical attacks and chemical insults (Haldane, 1929). Self-replication possibly came from the RNA World, where RNA molecules self-assemble and self-replicate out from amino acid building blocks (Joyce, 1989).
A site in Lenia can be thought of as a “molecule” (or a “concentration of molecules” in continuous case). Consequently a kernel would be a type of molecular force or chemical reaction, influencing surrounding molecules according to distance and concentration. Single-channel solitons, including those in the original Lenia, would resemble simple microscopic lifeforms (e.g. bacteria, archaea, viruses), possess self-organization, self-replication, symmetry, individuality, motility, etc.
Yet, to our knowledge, most of the “lifeforms” patterns that were discovered in Lenia were manually evolved from already stable SLP patterns (see here). Several key behaviors were missing in the original Lenia to push the comparisons further, such as pattern-emission. Interestingly, in less than 15000 training steps, IMGEP-HOLMES could discover many “lifeform” solitons that seem to arise from a “soup” of fluid-like patterns capable of pattern-emission, when their existence remained an open question in 2D Lenia. Examples discoveries are shown in the below videos.
These results confirm the importance for meta-diversity search in the context of automated discovery in morphogenetic systems. Indeed, the more interesting behaviors emerged out of an unexpected niche of “wavy” patterns, isolated in the bottom-right part of IMGEP-HOLMES tree hierarchy, whereas we would rather expect such structures to evolve from the already-stable SLPs isolated in the left branch of the hierarchy.
Complete Gallery of Discoveries
Below, we provide an interface to browse through all the patterns discovered by the different IMGEP variants implemented in the main paper. We can qualitatively observe that the discoveries of an IMGEP relying on a monolithic architecture for the behavioral embedding design tends to bias the final discoveries (both for hand-defined and unsupervisedly-learned features), and that therefore the final discoveries are unlikely to be aligned with the interests of a final end-user.
Filter
Notice: Due to storage restrictions, we provide the full database of discoveries for only the first repetition (out of 10 repetitions per IMGEP variant). All the exploration experiments start by 1000 runs of Random Exploration, this is why all the patterns numeroted from 1 to 1000 are the same for a given seed (repetition number).
Acknowledgements
We would like to thank Chris Reinke and Bert Chan for useful inputs and discussions for the paper, as well as Jonathan Grizou for valuable suggestions.